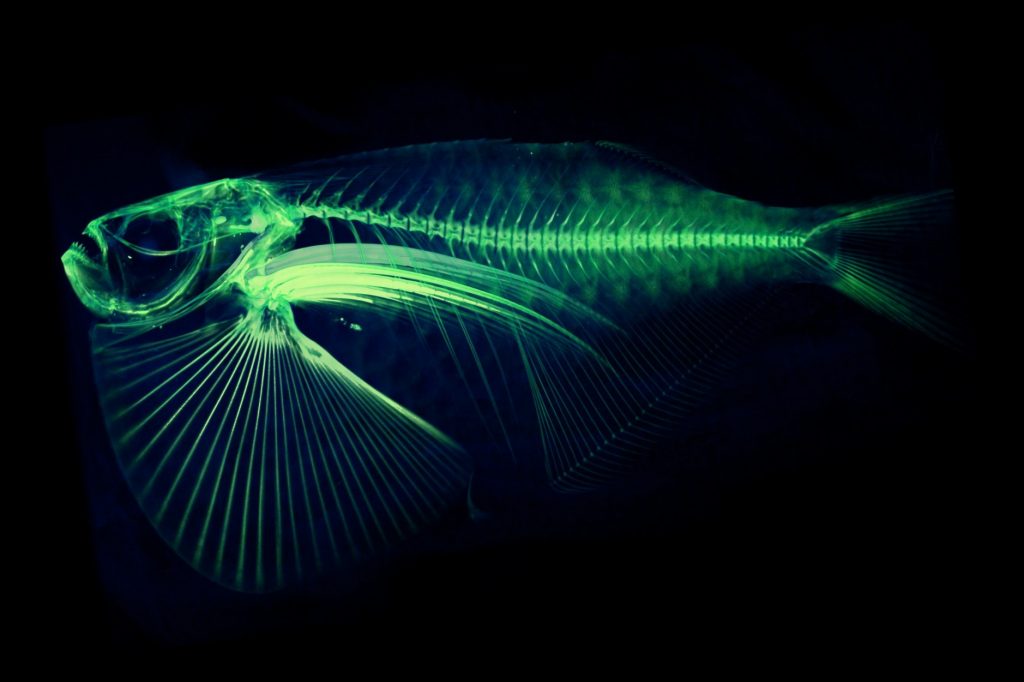
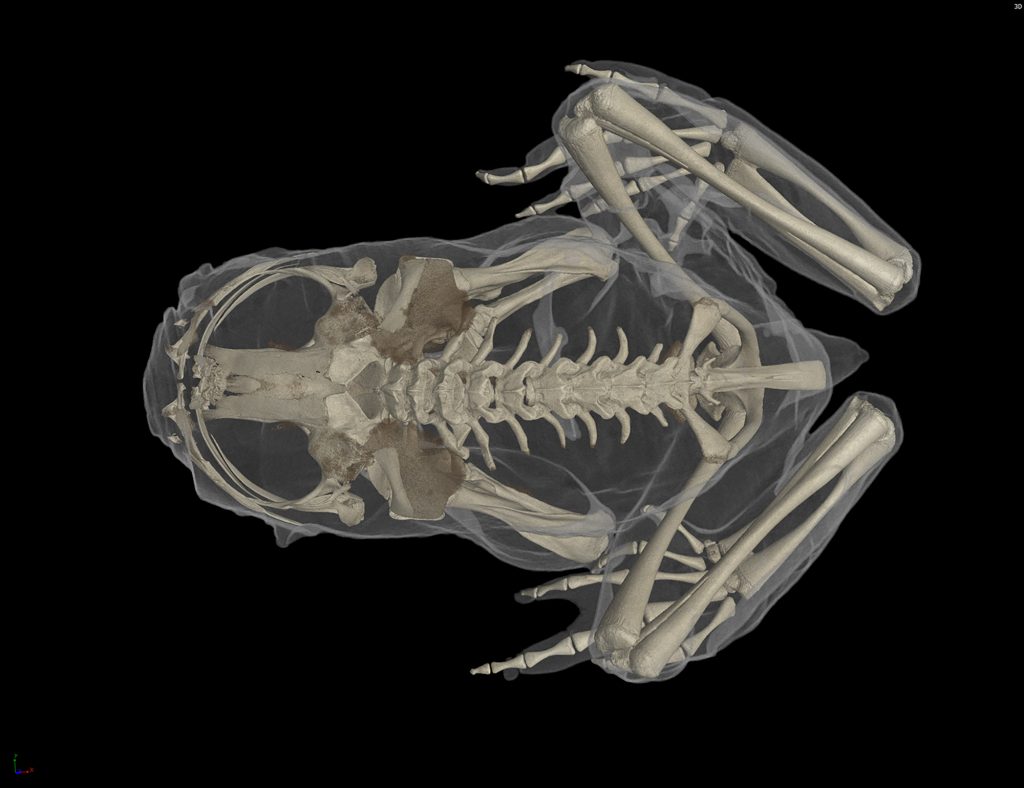
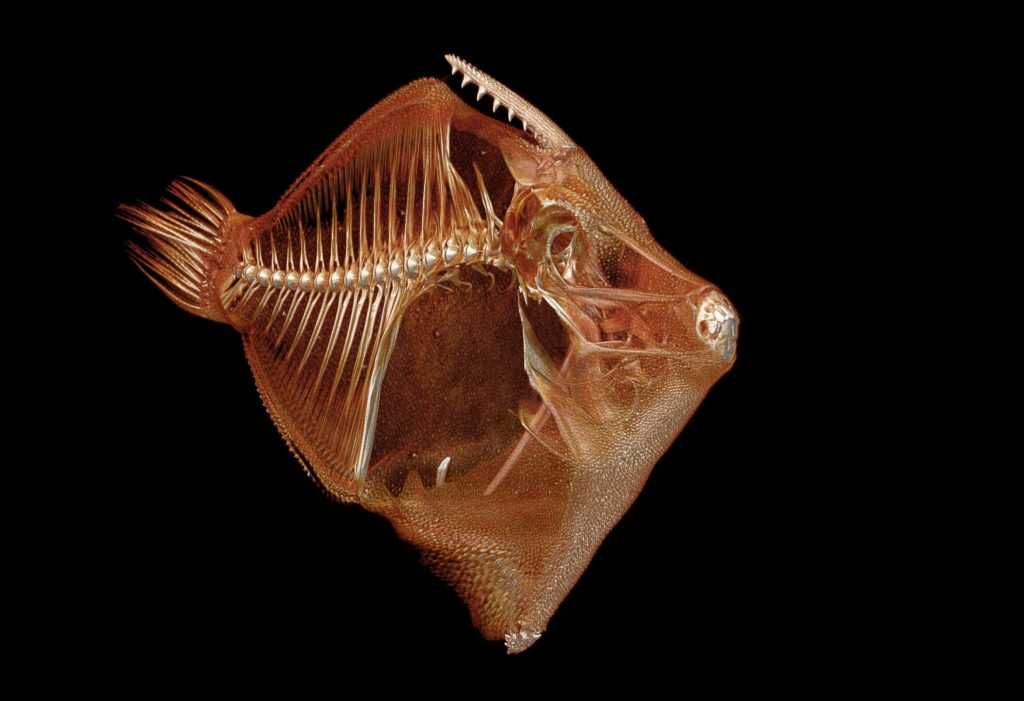
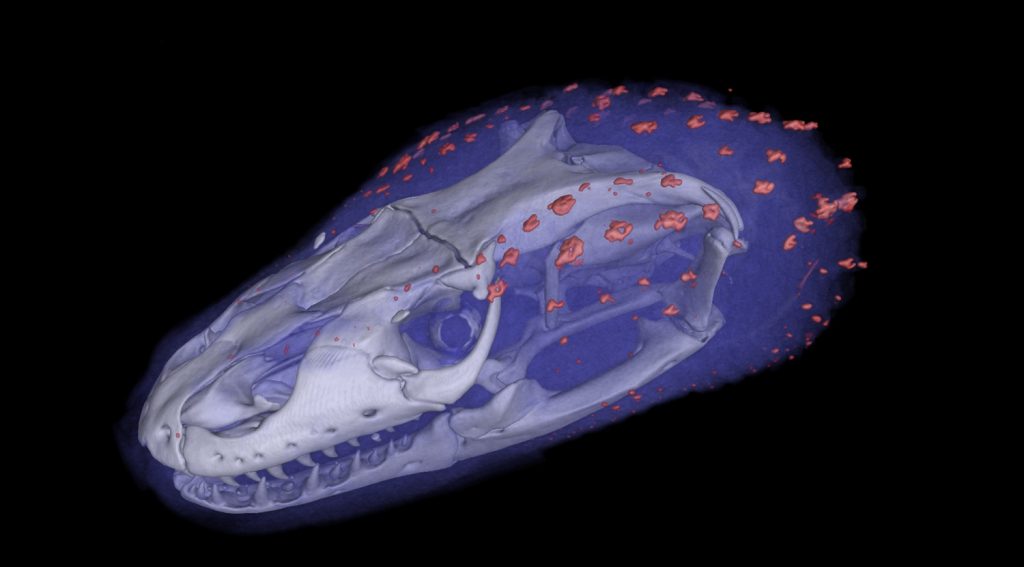
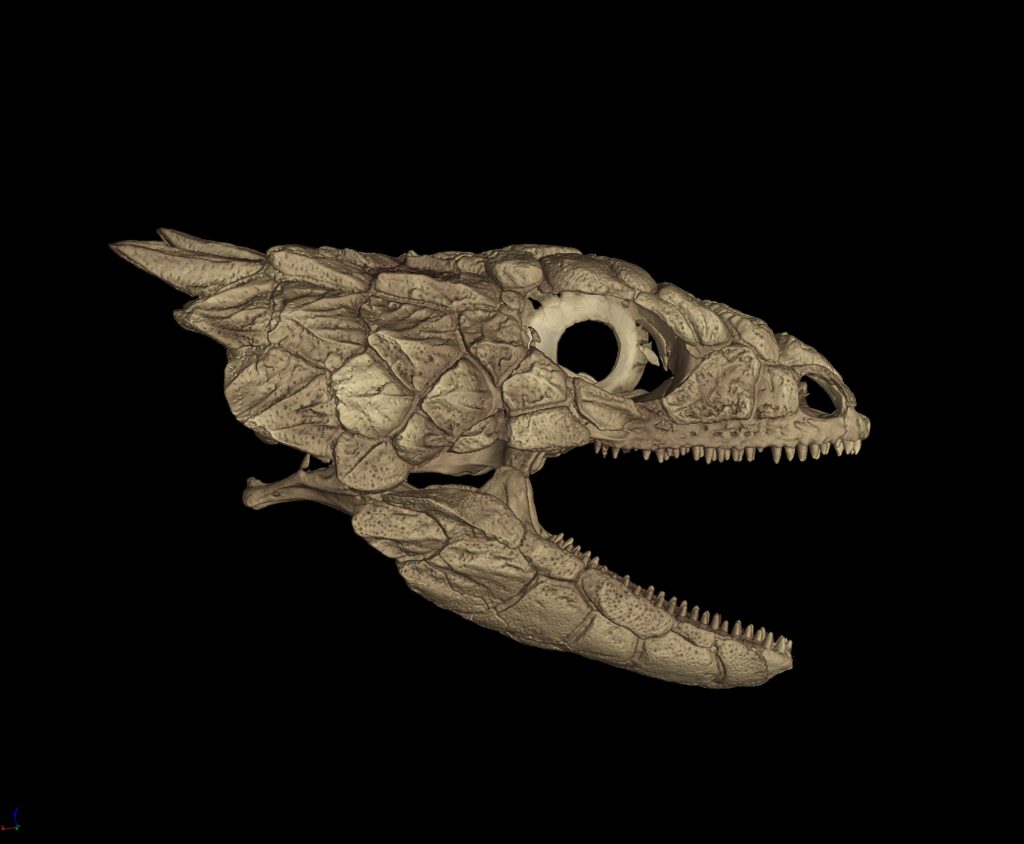
A $2.5 million National Science Foundation grant will launch a new initiative to “teleport” specimens from museum shelves to the internet by CT scanning 20,000 vertebrates and making these data-rich, 3-D images available to researchers, educators, students and the public.
The oVert project, short for openVertebrate, complements other NSF-sponsored museum digitization efforts, such as iDigBio, by adding a crucial component that has been difficult to capture — the internal anatomy of specimens.
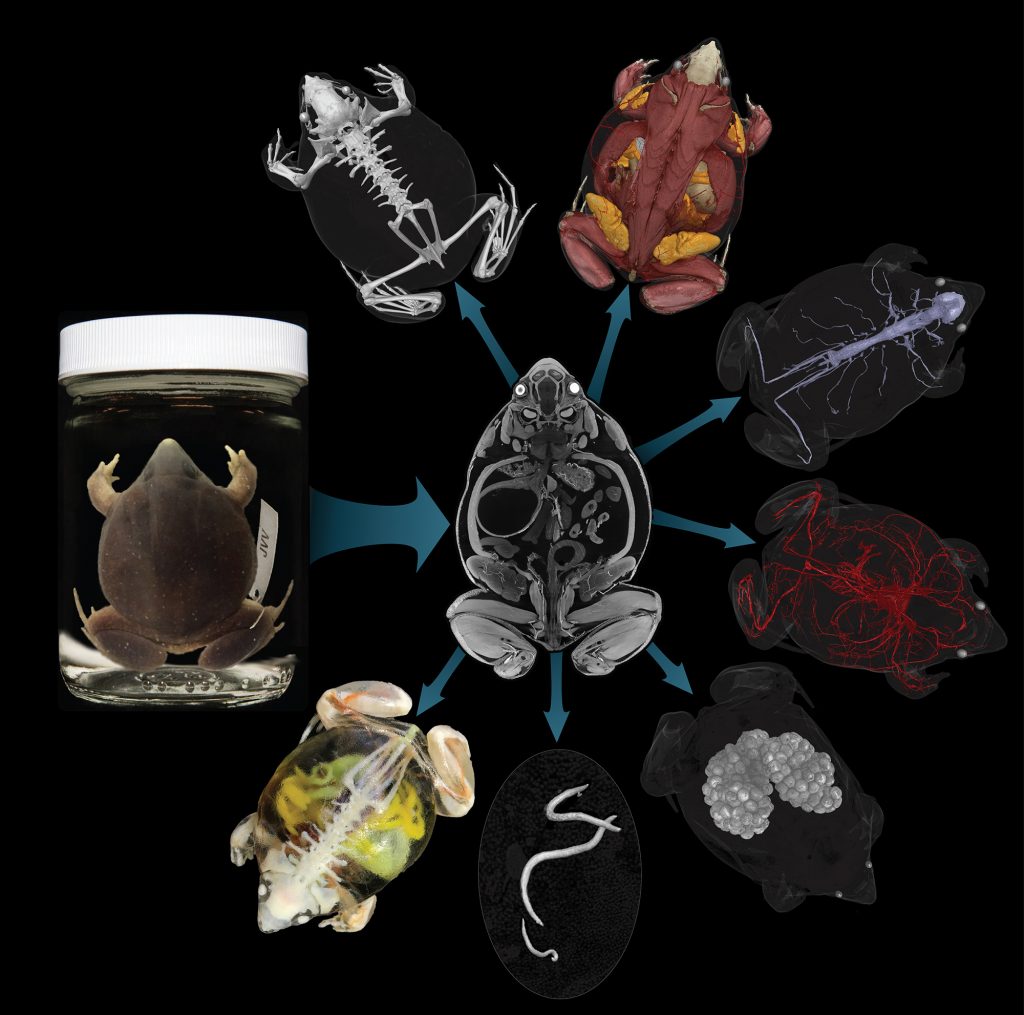
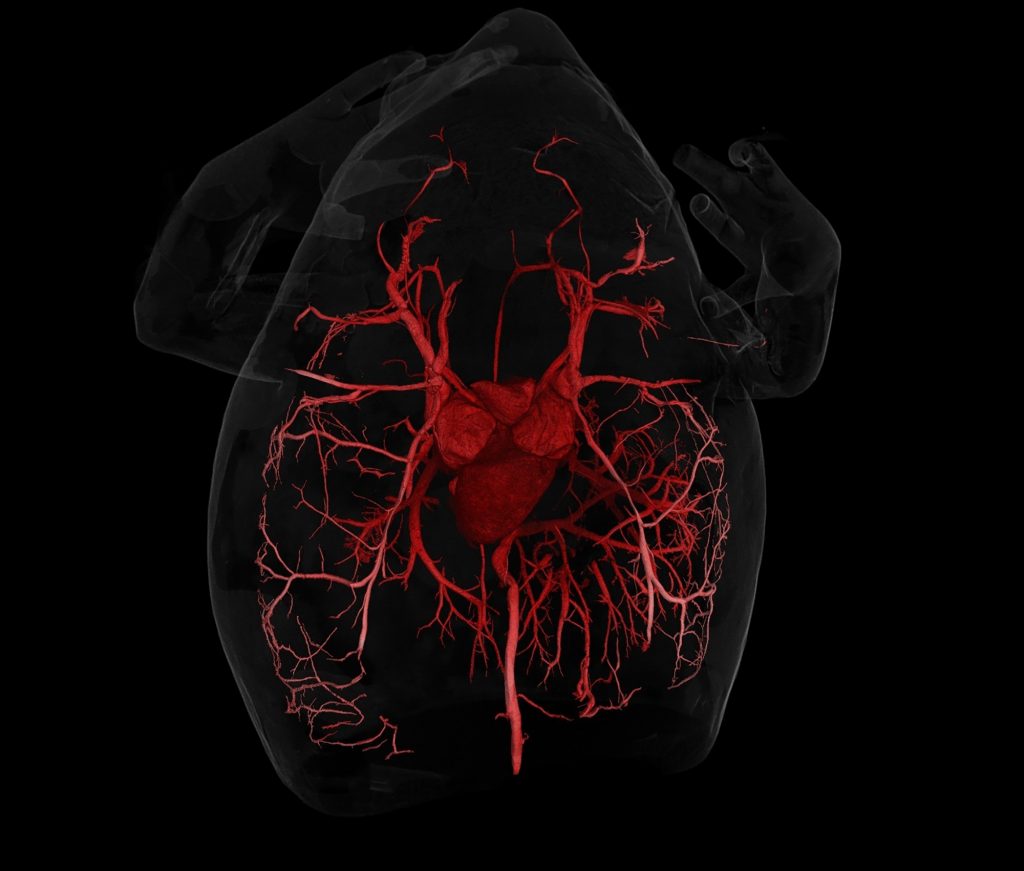
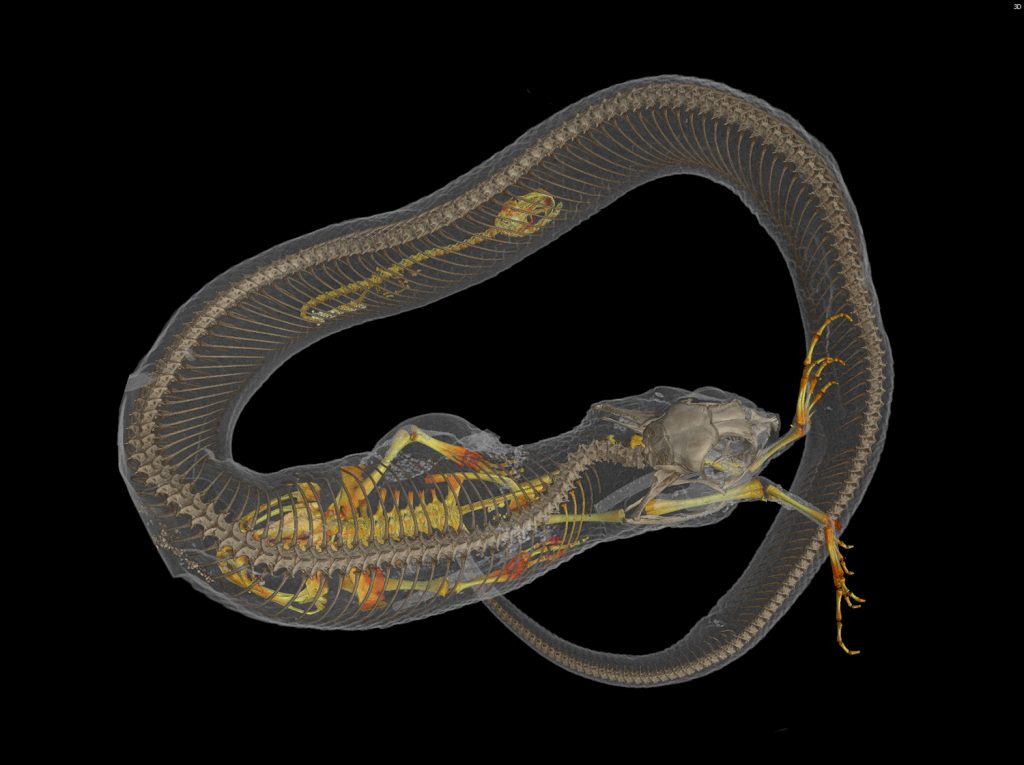
With virtual access to specimens, researchers could peel away the skin of a passenger pigeon to glimpse its circulatory system, a class of third-graders could determine a copperhead’s last meal, undergraduate students could 3-D print and compare skulls across a range of frog species and a veterinarian could prepare for a surgery on a giraffe at a zoo.
The project will encompass representative specimens from more than 80 percent of existing vertebrate genera, and a selection of these will also be scanned with contrast-enhancing stains to characterize soft tissues.
“Our goal is to provide data that offer a foothold into vertebrate anatomy across the Tree of Life,” said David Blackburn, the project’s lead principal investigator and associate curator of amphibians and reptiles at the Florida Museum of Natural History at the University of Florida, the grant’s lead institution. “This is a unique opportunity for museums to have a pretty big reach in terms of the audience that interacts with their collections. We believe oVert will be a transformative project for research and education related to vertebrate biology.”
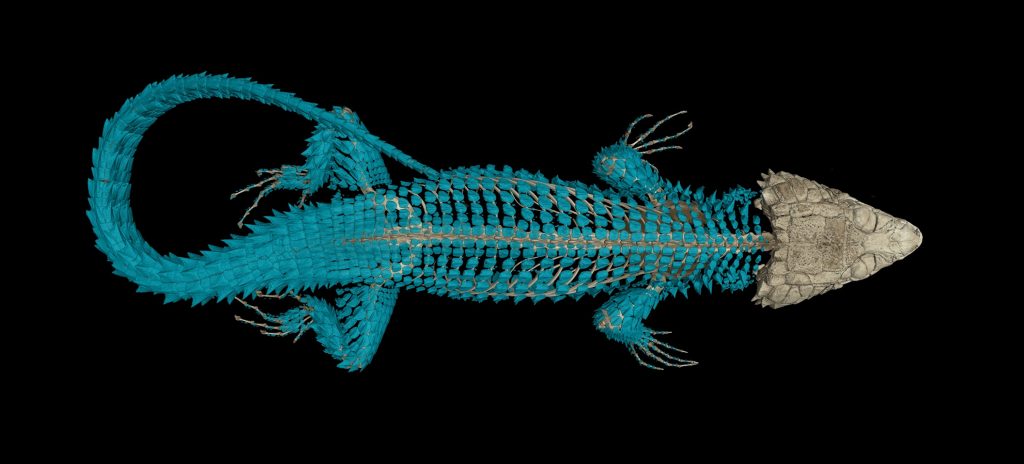
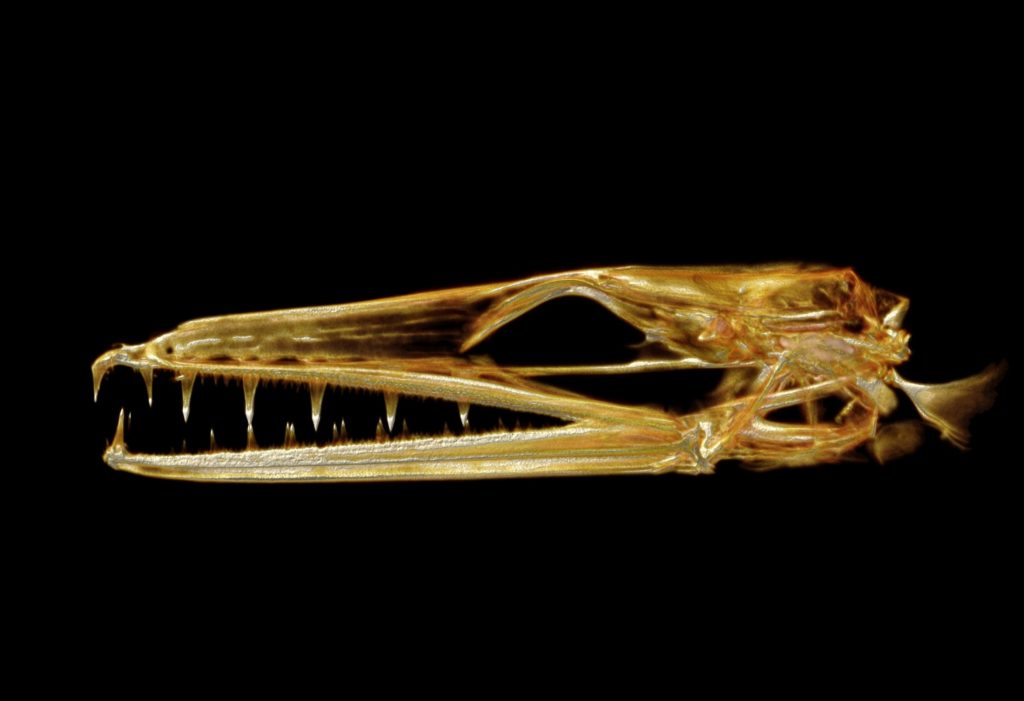
CT scanning is a nondestructive technology that bombards a specimen with X-rays from every angle, creating thousands of snapshots that a computer stitches together into a detailed 3-D visual replica. The image can be virtually dissected, layer by layer, to expose cross sections and internal structures. The scans allow scientists to view a specimen inside and out — its skeleton, muscles, internal organs, parasites, even its stomach contents — without touching a scalpel.
Images scanned for the project will be housed in MorphoSource, a public database created by Duke University that scientists, educators, students or the curious can mine for 3-D data on their species of interest.
“It’s exciting to glimpse the inside of rare and precious specimens, but that is just the beginning,” Blackburn said. “This opens up a whole world of vertebrate biology research.”
Advances in understanding the structure, function and evolution of genes and genomes have outpaced phenomics — the study of how genes interact with the environment to produce physical traits, or phenotypes. By providing a searchable digital encyclopedia of thousands of vertebrate phenotypes, oVert could be a valuable resource in narrowing the gap, Blackburn said.

“This is moving phenomes into big data science,” he said. “Even though traits have been the interface between development, paleontology, evolution and systematics since Aristotle, the big data we tend to think about are genomics and species distributions.”
CT scanning offers a wealth of data, but it’s expensive and time-intensive, limiting the number of specimens that can be scanned, Blackburn said. To maximize the project’s usefulness, researchers will select and scan “super specimens,” those that are representative of a species, are in good condition and have corresponding data in iDigBio, such as where, when and by whom they were collected. Ideally, the specimen would also have genetic data.
One of the key features of the initiative is making these data freely available, Blackburn said.
“We’re not going to hoard the data,” he said. “The realm of questions that can be asked in biology is often constrained by what is accessible. This lowers the barrier for people to work on a broader diversity of vertebrate life.”
In addition to increasing researchers’ access to vertebrate diversity, the oVert team also envisions the anatomical images being used in universities and schools, inspiring the next generation of scientists, engineers and mathematicians. The Florida Museum will partner with UF’s Center for Precollegiate Education and Training to develop and run yearly workshops with middle and high school science teachers to create lesson plans based on digital and 3-D printed specimens.
The project, which officially begins Sept. 1, is not without its challenges: Providing virtual access to specimens still requires boots-on-the-ground work. Specimens must be hand-selected, shipped, tracked, scanned, uploaded to MorphoSource and integrated with corresponding data in iDigBio. Multiply this process by 20,000 across 16 museum collections, and the workflow can quickly become daunting, Blackburn said.
“It’s not going to be without some growing pains.”
Sixteen research institutions are included in the grant: UF, the Academy of Natural Sciences of Drexel University, the California Academy of Sciences, Cornell University, the Field Museum of Natural History, Harvard University, Louisiana State University, the Scripps Institution of Oceanography at the University of California, San Diego, Texas A&M University, the University of California, Berkeley, the University of Kansas, the University of Michigan, the University of Texas, Austin, the University of Washington, the Virginia Institute of Marine Science and Yale University.
Duke University and the University of Chicago are also partners in the oVert project.
Source: David Blackburn, dblackburn@flmnh.ufl.edu, 352-273-1943
Learn more about oVert.
Learn more about Research and Collections at the Florida Museum.