
Photo by Jeff Gage
Nestled snugly at the end of Dickinson Hall’s west wing, in a room with bleached concrete floors, bleached counters and bleached walls, an ornithologist grinds the 2,000-year-old liquid-nitrogen drenched bones of a fossil rail—a type of bird—with a porcelain mortar and pestle until they yield a powdery dust. Jeremy Kirchman, a zoology doctoral student working in the Florida Museum of Natural History’s ornithology range, is extracting DNA fragments from the 500 to 2,000 year-old bones so he can answer questions about how these flightless birds evolved on the 12 Pacific Islands where they were collected over several decades by archaeologists.
Ancient DNA is a burgeoning field of study, and for museum curators and systematists—scientists who study how species are related evolutionarily—it offers a powerful tool for understanding how and when species originated, diverged and co-evolved.
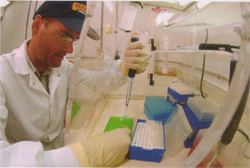
Photo by Jeff Gage
In the “ancient DNA lab” at the Florida Museum, Kirchman retrieved useful DNA strands from about 20 percent of the rail specimens. All of the viable samples were found in cave or rock shelter sites, protected from the elements. Which makes sense, given that the actions of sunlight and water act to destroy rapidly the nucleotide bonds of DNA. Nucleotides are chemical compounds that form the structural units of a DNA strand. Water and light decay them into bits and pieces too small and degraded to be useful to scientists.
“The amount of DNA you can get from a bone varies depending upon the condition of the specimen,” Kirchman said on a sunny August morning in the museum’s courtyard. “Dry, dark caves yield the best specimens.”
Finding useful DNA from the bones of an animal dead for thousands of years may at first glance appear to be a “needle-in-a-haystack” search. But amazingly, it happens with regularity. It has less to do with the specimen’s age than with its quality of preservation.
For a scientist trying to answer questions about how species within a family evolved, only a segment of DNA is needed, but it has to be the right one. Typically, a certain DNA segment will be targeted for study, recognizable by the order of the first few base pairs. It is these pairs that we recognize as the alphabet soup of DNA: A, C, T and G.

Photo by Jeff Gage
“You typically can only get sequences that are 100 to 250 base pairs long,” Kirchman said, because the DNA from fossil specimens is usually highly fragmented. But these hotly sought-after fragments can yield enough information to make conclusions or inferences about the relatedness of two or more species.
When an animal dies, the DNA molecule programming its cells begins to decay. But if protected from sun and water over hundreds or thousands of years, this decay can be slowed.
Of Lice and Birds
Some eclectic studies have passed through the doors of the ancient DNA lab at the Florida Museum of Natural History. When two pre-Columbian mummies were recovered from Peru, the researchers discovered that the disembodied heads-probably disturbed by looters-held hundreds of tiny, preserved head lice. A few lice were plucked and sent to Florida Museum assistant curator of mammalogy Dr. David Reed, who studies the co-evolution of lice and mammals. Reed enlisted Jeremy Kirchman’s expertise in ancient DNA extraction (see main story) to test a widely-accepted assumption about human lice evolution and distribution. Additional lice samples were sent to a second ancient DNA lab in Marseilles to independently verify the results.

There are three kinds of head lice in the world: types A, B and C. Type A is the most common and is found world-wide, but type C-which evolved earliest-is rare and found mostly in Africa. Type B was first found in Central and North America, but was later assumed to have arisen in the Old World and been introduced to the New World by European explorers and colonists.
By analyzing ancient DNA extracted from the mummified head lice, Reed and Kirchman identified it as the world-wide common lice, type-A. Since the mummies, and by extension the lice, were radiocarbon dated to roughly 1,025 years ago, or 981 C.E., this supports the idea that type-B lice arrived to the New World later, probably with European explorers.
“The interesting question now becomes ‘Why Europe?’ ” David Reed reflected of the findings. “What was going on in Europe’s history that would have allowed a distinctly different louse to emerge that is now restricted in distribution to the New World, Europe and Australia?”
Another project sought to identify a bird feather collected in Polk County, Florida in 1968 and believed to be from an ivory-billed woodpecker. Kirchman collaborated with ancient DNA researchers at the Smithsonian Institution, where he trained originally, to verify the feather’s source. However, feathers are notoriously bereft of DNA. Just as human hair only has genetic material at the follicle, feathers only have DNA at the shaft, where the feather anchors in the skin. The feather in the Florida Museum’s collection was sealed in a picture frame prior to its donation, decreasing the chances of preserving viable DNA. Neither Kirchman nor the Smithsonian scientists found any DNA in the shaft, which in a way adds to the bird’s elusive mystique.
The last reliable report of an ivory-billed woodpecker in Florida was in 1924, according to The Nature Conservancy’s Web site. In 2004, researchers announced that they had filmed an ivory-billed in a swampy bayou in eastern Arkansas, but two years of intensive searching since then has not produced definitive evidence of the magnificent bird. Many experts say the jury is still out on whether the ivory-billed woodpecker still graces old-growth forests of the Southeast.
“But even the cave specimens had only about a 50 percent success rate,” Kirchman said, referring to his study sites. “The ones from villages and open water sites never worked.” The success rate of extracting DNA from century-old or newer museum specimens skyrockets to about 99 percent, he said. This is because the specimens are kept in dark drawers, and the temperature and humidity are controlled.
Kirchman started the ancient DNA lab at the Florida Museum to conduct his doctoral research. A “modern” DNA lab, located across a tree-spotted plaza from the museum, housed the necessary DNA sequencer and thermocyclers. Kirchman secured grants and matching funds from two members of his Ph.D. advisory committee for the purchase of a centrifuge, an ultra-violet chamber and other equipment. By utilizing existing equipment in the modern DNA lab, he kept the ancient DNA lab’s start-up costs below $8,000.
How It Works
It takes a surprisingly small bit of smashed fossil bone to produce the DNA strands which hold the answers to some of our questions about evolution.
“It takes about two days of biochemistry to obtain 10 drops of liquid in a tube with highly degraded DNA inside,” Kirchman said. But if you have the right DNA sequence then those 10 drops can be milked for a gold mine of information.
First, Kirchman uses a Dremel to drill out a half-centimeter square bone sample about a quarter the size of your pinky fingernail. Next, he drops the bone chip into a ceramic mortar and adds liquid nitrogen. Frozen solid, the bone becomes like glass. It shatters when he uses the pestle to grind it into dust.
Next, he places the bone dust in a test tube and adds about 10 drops of buffers, enzymes and silica with a micro-pipette. The silica binds to the bone cell DNA, and the tubes are centrifuged to separate the bound silica-DNA from waste liquid and bone debris. The DNA is then extracted from the silica and is ready to be copied.
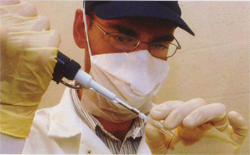
Photo by Jeff Gage
With his specimen now prepared, Kirchman caps the tubes and walks them to the modern DNA lab. Here, he uses a machine to activate an enzyme that begins a chain reaction, copying the targeted DNA sequence. Over three hours, it makes millions of copies from the original DNA fragment.
Kirchman now has his material but can not yet “read” the gene because the base pairs are too small, even with the most high-powered microscope. So he places the test tubes in a DNA-sequencing machine. The machine reads a color-coded fluorescing tag attached to each base pair of the sequence. These tagged pairs are then slid in a capillary tube past a laser beam. They fluoresce when passed in front of the laser’s light, and a photoreceptor records the color. A computer printout compiles the color sequences and translates it to an easily readable A-C-T-G format.
Voila. Two-thousand year-old bird bones come to light-a miniscule, but pivotal, portion of their genetic secrets laid bare.
The Clean Challenge
The biggest challenge in preparing the samples of ancient DNA for sequencing is preventing contamination with modern DNA. To prevent this, Kirchman showers before going to the lab and dons a clean lab coat, safety glasses, face mask and latex gloves before working. After each use, all surfaces of the lab are misted in a bleach solution and then meticulously wiped with sterile paper towels.
Kirchman said that in addition to these measures, he is careful about what he does. “You don’t want to eat a chicken sandwich for lunch then work up a bird sample, because then you have bird DNA all over your face,” he said.
Pipettes, mortar and pestle, and any other equipment used to prepare the sample are placed in an ultraviolet chamber and bathed in UV light for 12 minutes to kill any lingering DNA.
“The ancient DNA lab is essentially a clean room,” he said. “You want a DNA-free environment in which to prepare your sample.”
Kirchman used the DNA analyses to examine how quickly birds that colonize islands lose their ability for flight. He looked at the influence of geography and analyzed three genes to study how the flighted and flightless species were related to each other on the evolutionary tree of life. He expects to publish the findings in a peer-reviewed scientific journal next year.
Dr. Jeremy Kirchman received his Ph.D. in zoology this August from the University of Florida. He will be the new curator of ornithology at the New York State Museum in Albany.