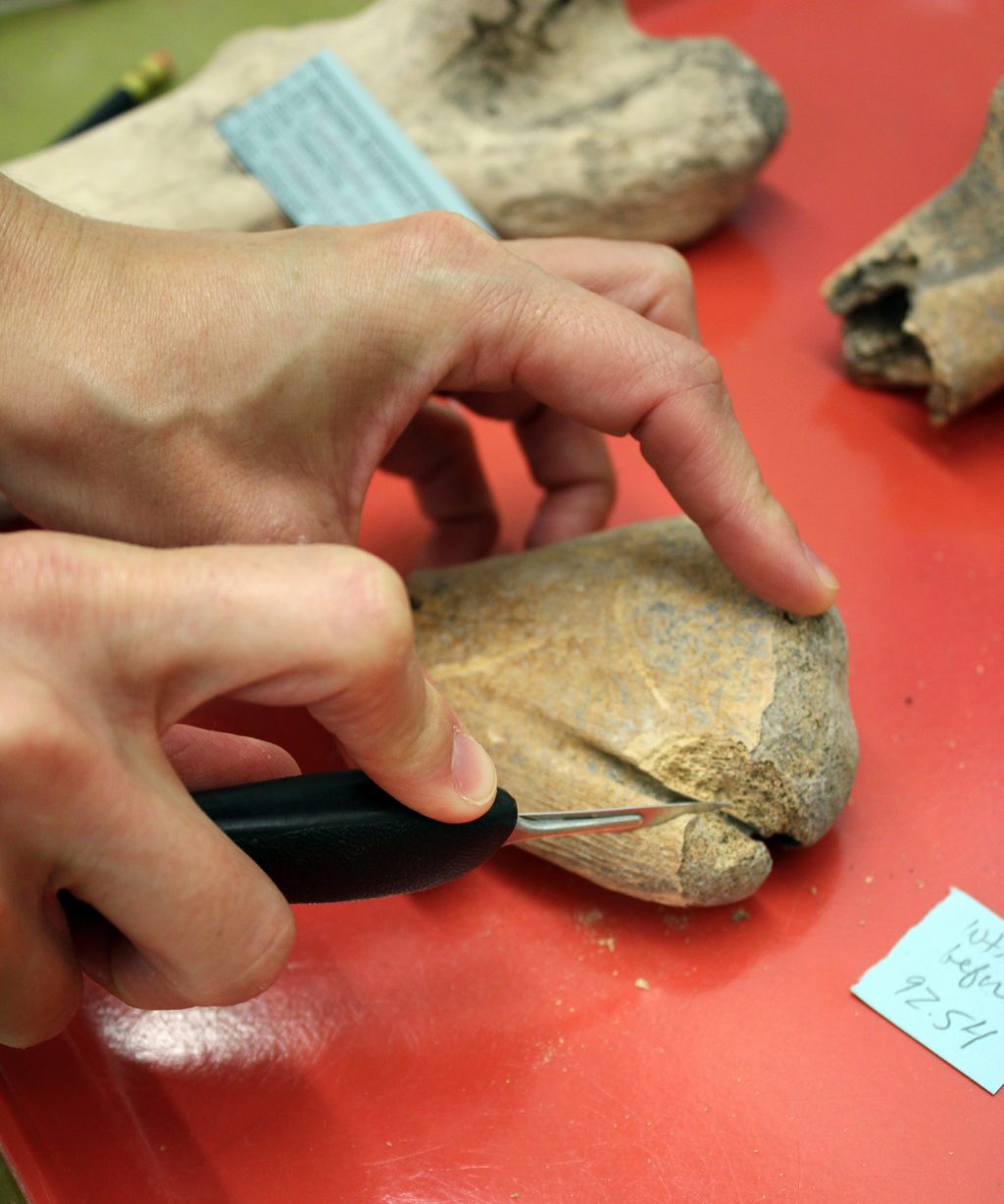
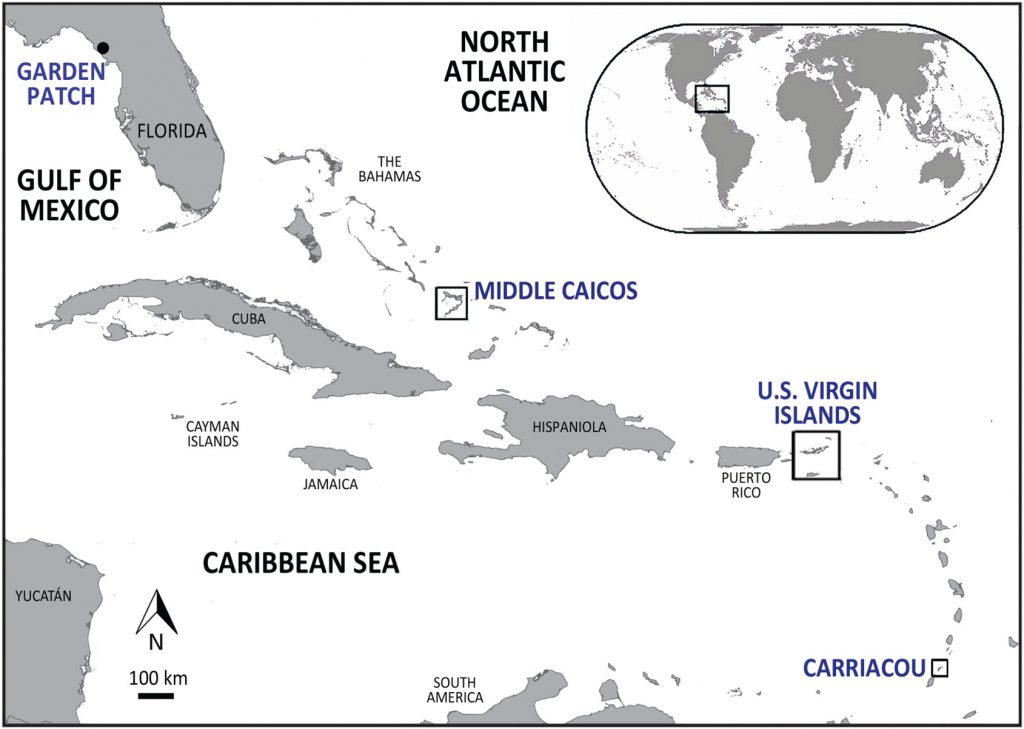
Thousands of years ago, the inhabitants of modern-day Florida and the Caribbean feasted on sea turtles, leaving behind bones that tell tales of ancient diets and the ocean’s past.
An international team of scientists used cutting-edge technology to analyze proteins from these bones to help identify which turtle species people fished from the ocean millennia ago. This can aid modern conservation efforts by helping construct historical baselines for turtle populations, many of which are now endangered,and illuminate long-term trends of human impacts.
The technique, known as collagen fingerprinting, allows scientists to visualize distinct chemical signatures in collagen, the main structural protein in bone, that are often species-specific. This provides a complementary alternative to comparing specimens’ physical characteristics and analyzing ancient DNA, two methods that can be unsuccessful for species identification in fragmented archaeological bones found in the tropics.
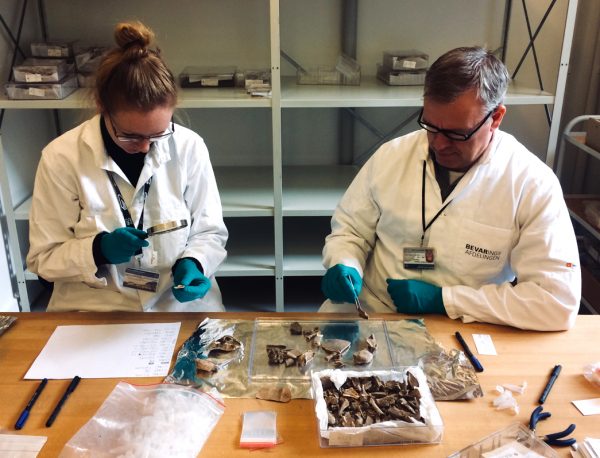

Photo courtesy of the National Museum of Denmark’s department of modern history and world cultures
Applying collagen fingerprinting to more than 100 turtle samples from archaeological sites up to 2,500 years old, the researchers found that 63% of the collagen-containing bones belonged to green turtles, Chelonia mydas, with smaller numbers of hawksbill turtles, Eretmochelys imbricata, and ridley turtles, Lepidochelys species. Some specimens previously identified as sea turtles from their skeletal features were in fact bones from snapping turtles, terrapins and tortoises.
“This is the first time anyone has obtained species-level information using proteins preserved in archaeological sea turtle bone,” said Virginia Harvey, the study’s lead author and a doctoral researcher in marine biology and zooarchaeology at the University of Manchester. “Our method has allowed us to unlock ancient data otherwise lost in time to see which species of turtle humans were targeting thousands of years ago in the Caribbean and Florida regions.”
Globally, sea turtles have been exploited for millennia for their meat, eggs, shells and other products. Today, they face threats from habitat loss and disturbance, poaching, pollution, climate change and fisheries. Only seven species of sea turtle remain, six of which are classified as vulnerable, endangered or critically endangered. Gaining a historical perspective on how turtle populations have changed through time is a crucial component of conserving them, Harvey said.
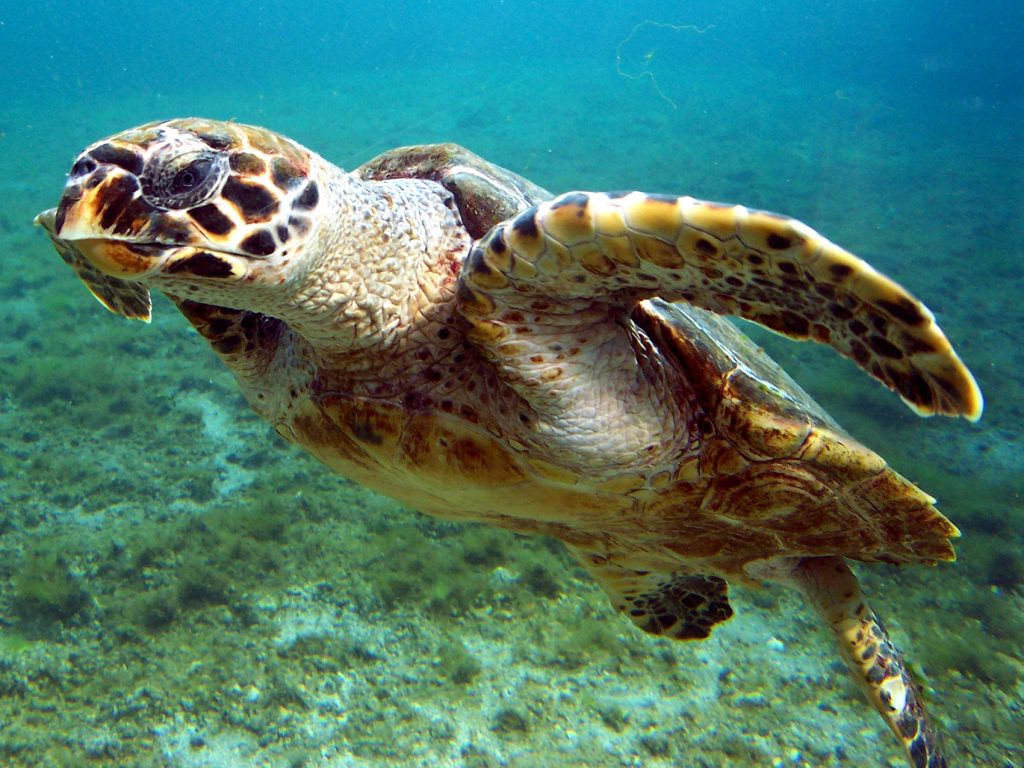
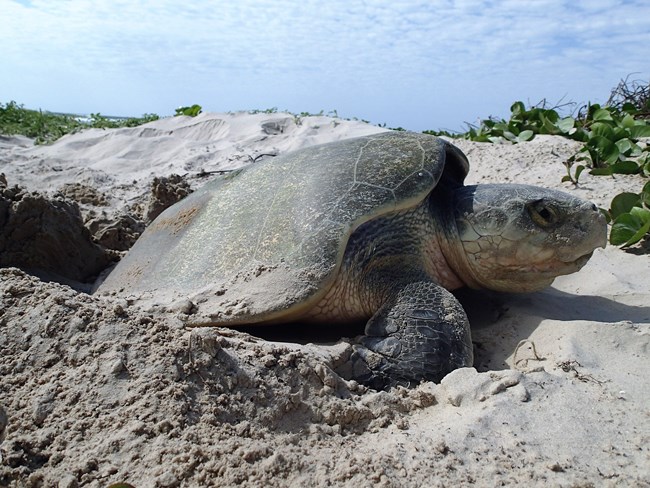


One of the research team’s initial goals was to discern whether any collagen still survived in ancient turtle bone remains. In an analysis of 130 archaeological turtle samples, the team was able to detect collagen in 88%.
“We were very impressed with the levels of protein preservation in the turtle bones, some of which are thought to be up to 2,500 years old,” said study co-author Michelle LeFebvre, assistant curator of South Florida archaeology and ethnography at the Florida Museum of Natural History. “The fact we were then able to use the protein signatures for species identification to better understand these archaeological sites was very exciting.”
The team uncovered an unusual chemical signature in a small number of bone samples that could suggest they belong to a different species than those present in oceans today. But when the researchers attempted ancient DNA analysis on them, they found the material was too degraded.
“Where DNA sequencing can often give more accurate information about species identity, this molecule is very fragile and does not always survive too well in ancient samples from hot, humid climates,” said study co-author Konstantina Drosou, ancient DNA specialist at the University of Manchester.
In contrast, proteins are present in much higher concentrations and therefore more likely to survive in the archaeological record, Drosou said.
“Proteins are very sturdy molecules,” Harvey added. “The absence of preserved DNA in these samples means we have not been able to verify whether they represent a new species of sea turtle or not, but it does show us that our work here is far from complete. There is so much that we can still learn from the turtle remains at these sites and beyond.”
Using collagen fingerprinting to correct misidentifications based on physical characteristics was “a nice additional outcome of the study,” said Michael Buckley, senior author of the study and senior research fellow at the University of Manchester.
Susan deFrance, study co-author and professor in the University of Florida department of anthropology, said juvenile sea turtles are often misidentified because they are small and may lack the characteristics used to distinguish adult sea turtle bones.
“This is the first time we have been able to look so specifically into the preferred food choices of the site occupants,” she said. “At the Florida Gulf Coast site, they captured a lot of juvenile turtles. The positive species-level identifications of these samples could not have been accomplished without this collagen fingerprinting technology.”
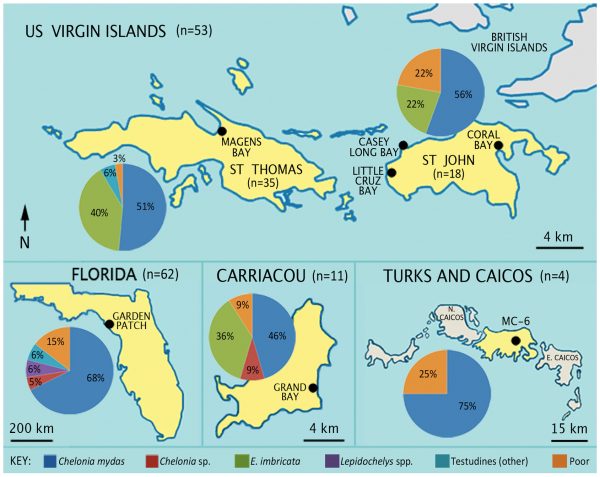
Image by Harvey et al. in Royal Society Open Science
At the same site, researchers found green turtle remains in both refuse heaps and mounds, but ridley turtle specimens were only found in mounds, suggesting they may have been reserved for feasting rituals, LeFebvre said.
“We knew these ancient people were eating sea turtles, but now we can begin to hone in on which turtles they were eating at particular times,” she said. “It’s no different than today – we associate certain foods with certain events. It’s how humans roll.”
The researchers are also eager to continue to apply collagen fingerprinting to other archaeological museum specimens, many of which have yet to be positively identified to the species level.
Harvey said she hopes the study inspires further research on sea turtles and other vulnerable and endangered animals.
“Now that this method is available, we hope that biologists, archaeologists and conservationists globally will continue this important work.”
The researchers published their findings in Royal Society Open Science.
Casper Toftgaard of the University of Copenhagen and Andrew Kitchener of National Museums Scotland, Edinburgh also co-authored the study.
Funding for the research came from the University of Manchester and the Royal Society.
Sources: Virginia Harvey, virginia.harvey@manchester.ac.uk;
Michelle LeFebvre, mlefebvre@flmnh.ufl.edu, 352-273-1917;
Konstantina Drosou, konstantina.drosou@manchester.ac.uk;
Michael Buckley, m.buckley@manchester.ac.uk;
Susan deFrance, sdef@ufl.edu